RNA degradation in yeast preprint
Deadenylation and RNA degradation: a "chicken and egg" problem
Our most recent results are now available as a preprint on bioRxiv. In short, it shows that, contrary to expectations, altering the speed of deadenylation form most yeast mRNA is not likely to change their half-life. Thus, the short mRNA poly(A) tails observed for stable mRNA (Lima et al., 2017) are potentially a consequence of mRNA stability. Stable RNAs have only more time to be targeted by deadenylation. Even if deadenylation is required to trigger mRNA degradation, previous modeling of the process has shown that, to fit experimental data, the oligoadenylated version of stable mRNA needs to be degraded with slower rates than in the case of the unstable ones. The difference can vary by a factor of 17 in yeast to a factor of 1000 in mammalian cells (Parker and Bartel labs).
I like to think of this situation as a "chicken and egg" problem. While the prevalent view is that deadenylation is a pre-requisite for mRNA degradation, the funny thing is that one can look at experimental results the other way around. What if deadenylation is in fact not required for mRNA degradation and occur in parallel to it ?
While our results are presented on biorxiv as "Contradictory", they are very much in agreement with previously published data, although a reinterpretation of what these previous publications show is required. I will only give here a simple example, from a beautiful paper published more than 30 years ago by David Herrick, Roy Parker and Allan Jacobson. In that paper, the authors looked at mRNA decay rates using a thermosensitive version of one of the subunits of the RNA polymerase II. While the change in temperature used might affect the measured half-lives, some of the presented experiments are very intriguing. They probably show, for a poly(A) selected population, the extent of deadenylation at different time points:
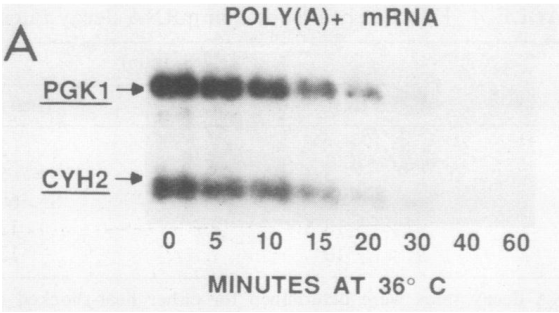
The decrease of mRNA levels in the figure shown left (or above) could be interpreted as a fast degradation of the poly(A) species. A simpler explanation for this observation is that what we are actually witnessing is the disappearance of the long poly(A) mRNA species. Thus, the experiment is probably estimating the deadenylation rate for PGK1 and CYH2, two stable transcripts.
I am not sure how easy it is to separate the poly(A) minus fraction of mRNAs, but, if this separation worked well, the figure at the right (or below) shows that somewhat oligoadenylates species of PGK1 and CYH2 are quite stable. Thus, it seems that deadenylation and mRNA degradation are not particularly linked in these experiments.
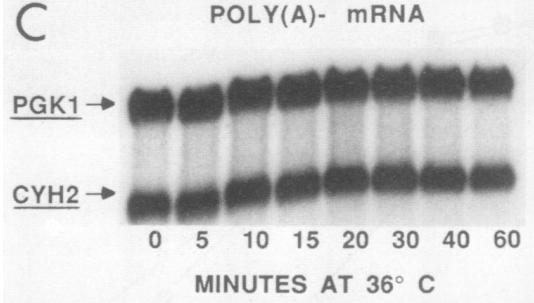
Not shown here is the situation of an unstable mRNA, in which deadenylation and degradation have probably similar speeds. In this situation, estimates of deadenylation rates can be particularly comple, unless one uses a mathematical model and as many experimental points as possible.
If our results indicate that deadenylation rates are unlikely to dictate RNA degradation speed, this does not mean that regulation of deadenylation cannot affect the levels of an mRNA, its localization or its translation. On the contrary, it allows, superimposed on a background of variable deadenylation rates,