M2 student position for 2024 (now filled)
M2 training period position to study mRNA degradation (now filled)
A position to study the biochemistry and molecular biology of RNA degradation in the pathogenic yeast Cryptococcus neoformans is available starting January 2024. Compared to Saccharomyces cerevisiae, a fantastic model for understanding mRNA degradation, C. neoformans retained splicing-related proteins, such as those composing the core exon-exon junction complex. Moreover, unlike in budding yeast, the presence of introns is an absolute requirement for gene expression. Finally, recent technical developments allow genetic manipulations of cryptococcal strains that was not possible until now.
Outline of the project and its context
How eukaryotic cells resist viral infections ? How mRNA degradation mechanisms regulate gene expression and suppress harmful transcripts ? How degradation pathways choose the “bad” transcripts that need to be rapidly degraded ? Why and how mRNA translation can both lead to stabilization and destabilization of mRNA ?
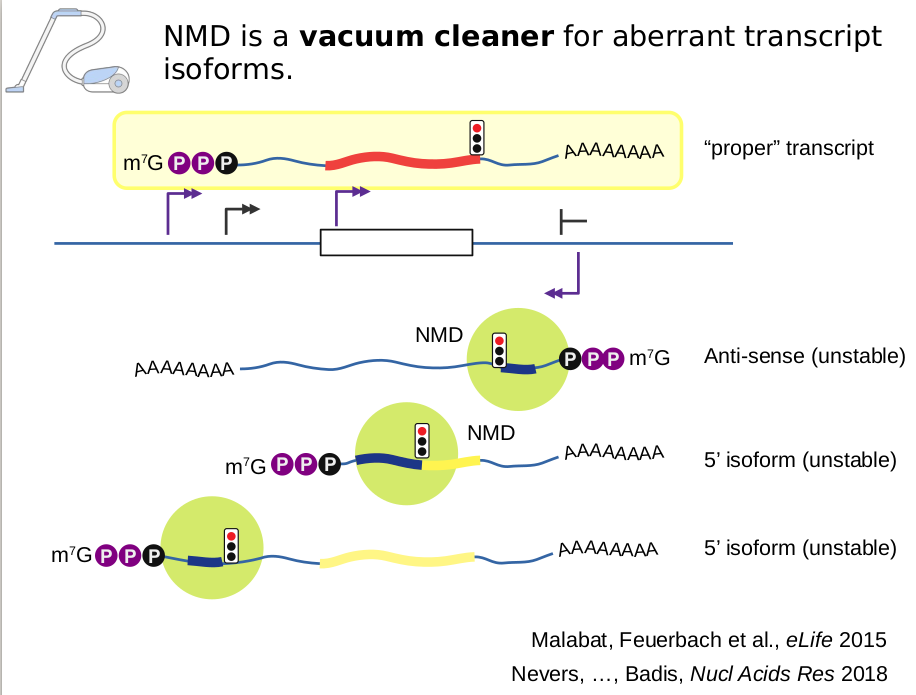
We propose to answer these questions by focusing on NMD, short for nonsense-mediated mRNA decay. In addition to viral control, NMD serves to regulate specific transcript levels and to keep at bay large amounts of transcripts that are by-products of RNA polymerase activity. NMD is triggered by a translating ribosome that encounters an aberrant stop codon and the study of NMD mechanisms led to major questions, such as: a) how the NMD machinery is recruited to specific transcripts? and b) how recruitment of NMD factors is linked with mRNA degradation ?
Our group began answering these question by employing large-scale affinity purifications and genetic screens using the budding yeast Saccharomyces cerevisiae as a model organism (EMBOJ 2018, Nucl Acids Res, 2021). We identified NMD factors that escaped previous investigations and were able to show that yeast employs NMD machineries that have equivalents in many other organisms including humans. Our preliminary data on NMD complexes support the hypothesis that regulated mRNA degradation plays a major role in stress adaptation. Targeted mRNA degradation and its regulation during stress might represent an important virulence factor for fungal pathogens, such as Cryptococcus neoformans, a remote cousin of budding yeast separated of it by about 1 billion years of evolution. We propose to investigate NMD complexes and their function in this organism and perform comparative functional analyses to decipher molecular mechanisms of mRNA degradation that are either specific to branches of eukaryotic evolution or are universally conserved. These investigations will involve affinity purification of RNA-protein complexes, quantitative mass-spectrometry and RNA sequencing. Knowledge of these mechanisms can be useful in understanding the pathogeny of fungal infections, in developing new therapeutic strategies and in better understanding the molecular biology of gene expression in living cells.