Publications
All the publications:
 PubMed search for C. Saveanu →, for papers as first author→ or last author→ (links open in new tab).
PubMed search for C. Saveanu →, for papers as first author→ or last author→ (links open in new tab).
Selected publications (or preprints)
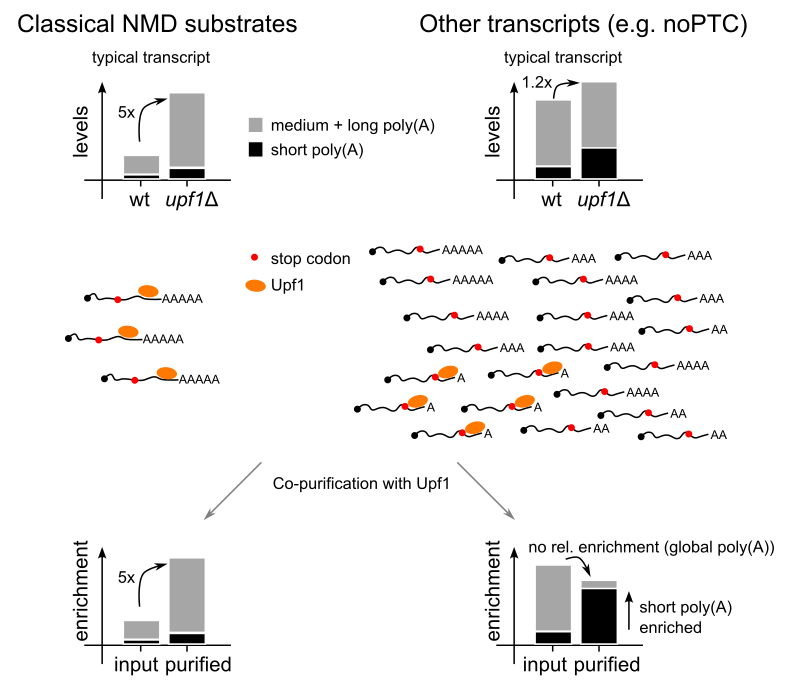
Gouhier T, Saveanu C*, Badis G* (2026) Nonsense-mediated mRNA decay factors target short poly(A)-tailed mRNAs lacking a premature termination codon. Nat Commun. PMID: 41997935 (pre-print on HAL-Pasteur)
Nonsense-mediated mRNA decay was always seen as a peripheral RNA degradation pathway, dealing with defective or unwanted mRNA. We might have discovered a new class of such mRNA that, instead of being separate from, is in reality part of most mRNA. The main feature of this new class of NMD substrates is the presence of a short poly(A) tail. Direct RNA sequencing results using Nanopore allowed to show that Upf1, a core NMD factor, binds specifically to short poly(A) mRNA for thousands of different transcripts in yeast. This binding was not only specific but, for a few tested transcripts, led an acceleration of degradation of the short poly(A) fraction. The mechanism by which Upf1 recognizes specifically the subset of mRNA with short poly(A) versus the same mRNA with longer poly(A) tails remains a mystery.
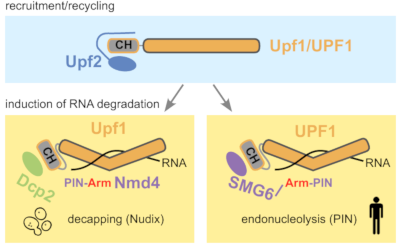
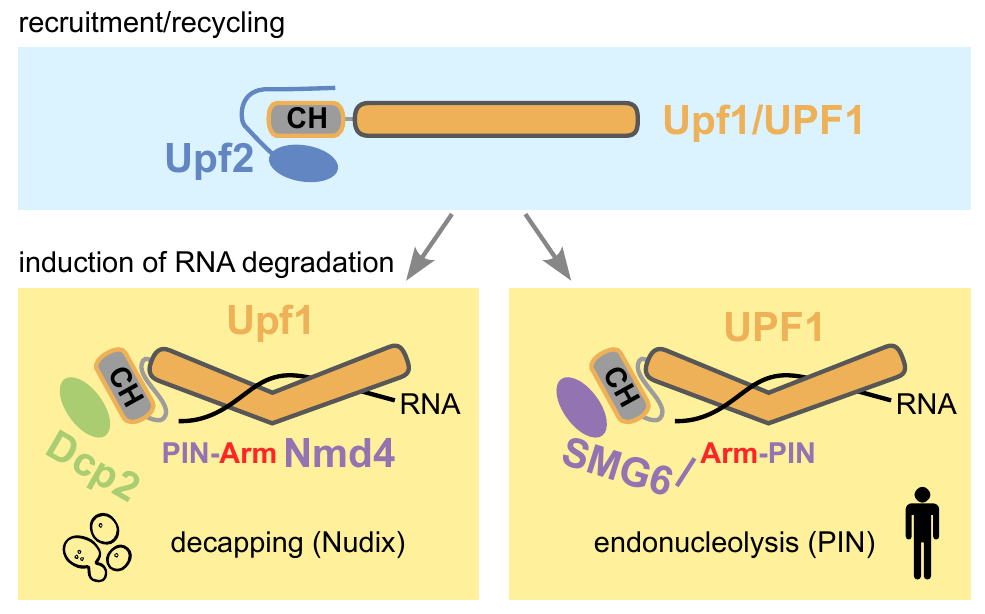
Ruiz-Gutierrez N, Graille M, Le Hir H, Saveanu C* (2026) Biochemical insights into the conserved interactions of NMD factors from budding yeast to humans. J Mol Biol. review PMID: 41819420 (pre-print on HAL-Pasteur)
Ever wondered to what extent protein-protein interactions are rewired during evolution ? In collaboration with the laboratories of Hervé le Hir and Marc Graille, we investigated old and new data about molecular interactions in NMD, focusing on knowledge obtained with the humble budding yeast, S. cerevisiae. One of the most interesting conclusions of this literature review was that the endonuclease SMG6, required for cutting NMD substrates in mammals, drastically diminished in size to the small Nmd4 protein in yeast. Moreover, its endonucleolytic activity was replaced by decapping, while the interactions of SMG6 with UPF1 were preserved in the form of the couple Dcp2/Nmd4 in yeast. This is a remarkable conservation over billions of years of evolution. It demonstrates also the importance of using a variety of organisms when studying complex molecular processes, such as NMD.
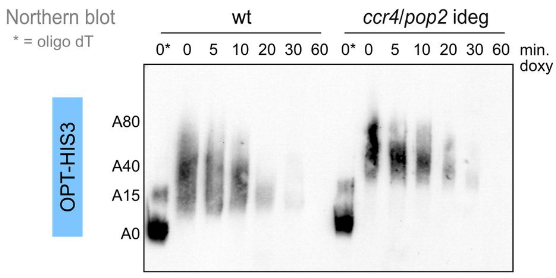
Audebert L, Feuerbach F, Zedan M, Schürch AP, Decourty L, Namane A, Permal E, Weis K, Badis G, Saveanu C* (2024) RNA degradation triggered by decapping is largely independent of initial deadenylation. EMBO J.PMID: 39233754
Biology manuals describe mRNA degradation as following an obligatory 1) deadenylation and 2) decapping sequence of molecular events. Careful examination of this assertion led to a different view about the life of mRNA in S. cerevisiae. For most unstable mRNAs, it turns out that deadenylation is not required for initiating mRNA degradation. However, we also identified situations in which deadenylation, rather than decapping plays a role in mRNA degradation. This work has been done in collaboration with the group of Karsten Weis at ETH Zürich, where the SLAM-Seq results have been obtained.
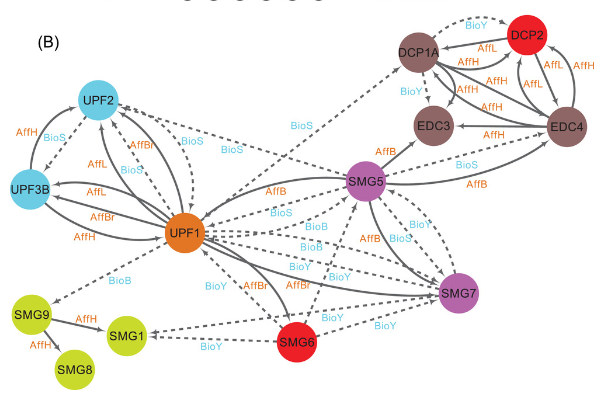
Gilbert A & Saveanu C* (2022) Unusual SMG suspects recruit degradation enzymes in nonsense-mediated mRNA decay. Bioessays e2100296. - review PMID: 35266563
In this review, we discuss the challenges in understanding mRNA degradation. Among them, the heterogeneity of the molecules of RNA, which are a mix of old and new, polyadenylated and deadenylated, full length and fragments. We also take a look at the conservation of the factors involved. Our previous work and recent collaborations with H. Le Hir's and M. Graille's laboratory clearly identified the equivalence of yeast and metazoan NMD co-factors. As an example, NMD4 retained the PIN domain typical of SMG-6 together with a small domain of interaction with Upf1. A specific SMG-6 motif identified by the work of the Graille team is important for the interaction of human UPF1 with SMG-6.
Namane A & Saveanu C* (2022) Composition and dynamics of protein complexes measured by quantitative mass spectrometry of affinity purified samples. book chapter in Methods in Molecular Biology 2477:225-236, Yeast functional genomics, (Springer), preprint at HAL pasteur-03384570v1 . PMID: 35524120 ISBN: 978-1-0716-2257-5
This methods article describes in detail the use of the ZZ motif derived from the S. aureus protein A to purify protein complexes from yeast (S. cerevisiae). It also includes the recipes for the preparation of IgG-magnetic beads and hints about mass spectrometry results analysis.
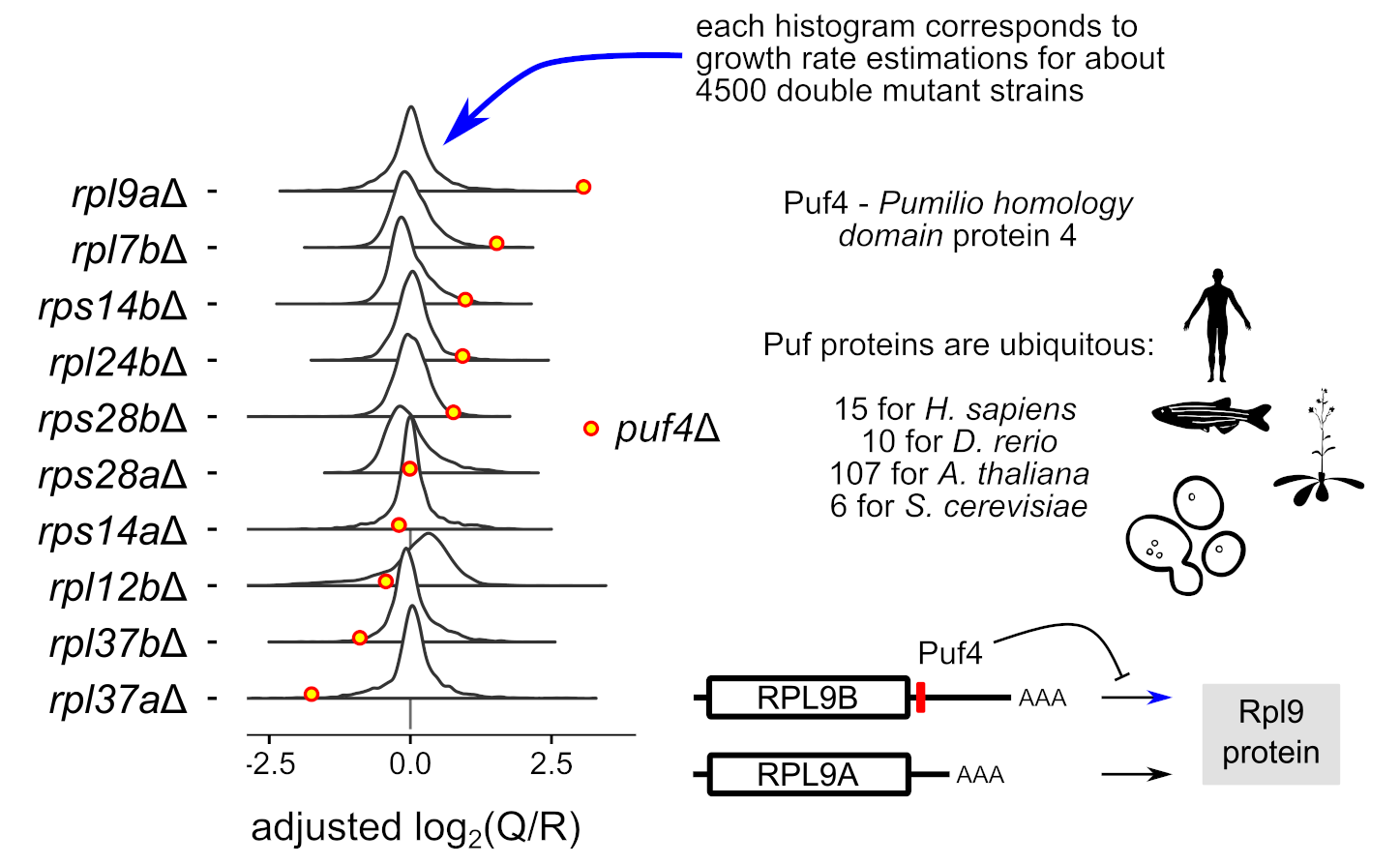
Decourty L, Malabat C, Frachon E, Jacquier A & Saveanu C*. Investigation of RNA metabolism through large-scale genetic interaction profiling in yeast. Nucleic Acids Research (2021), Sep 7;49(15):8535-8555. PMID: 34358317
The results can be explored through an interactive web interface at : cosminribo.shinyapps.io/GIM_interactions/→
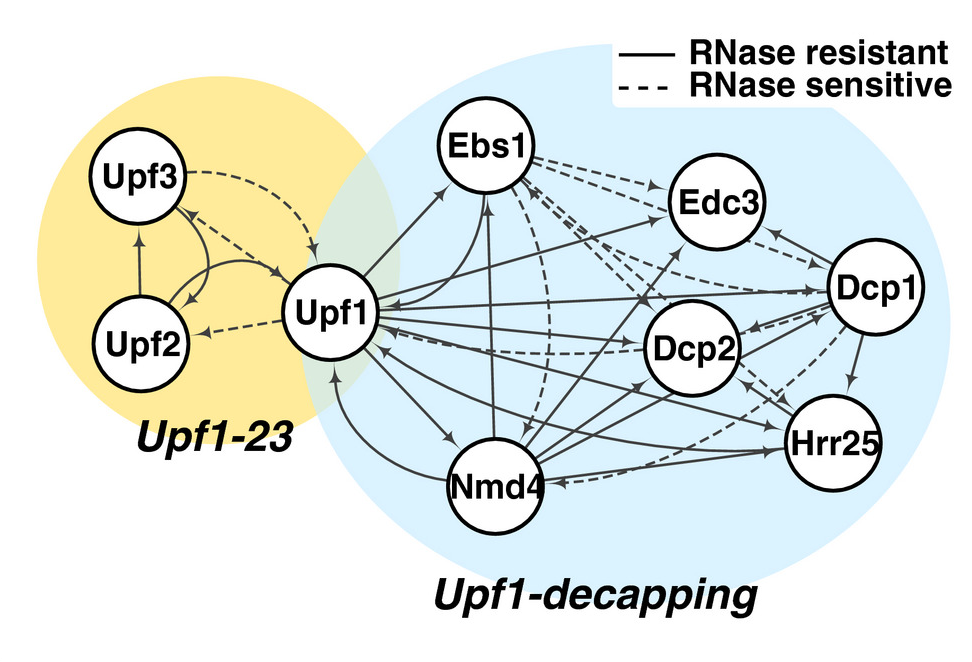
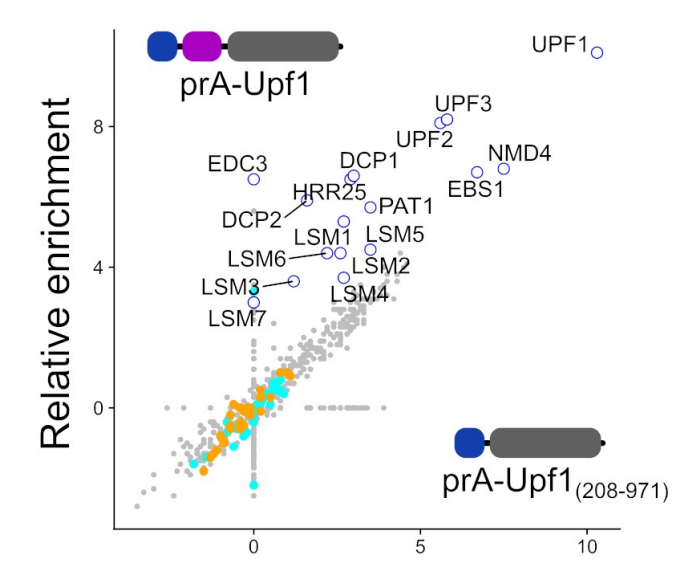
Dehecq M, Decourty L, Namane A, Proux C, Kanaan J, Le Hir H, Jacquier A & Saveanu C*. Nonsense-mediated mRNA decay involves two distinct Upf1-bound complexes. EMBO J (2018), Nov 2;37(21):e99278. PMID: 30275269
The protein association results can be explored through an interactive web interface at : cosminribo.shinyapps.io/NMD_complexes/→
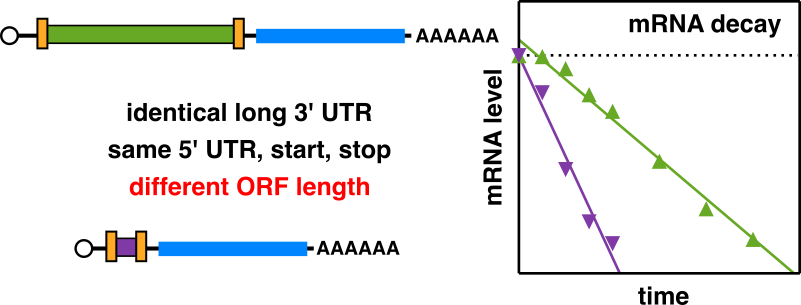
Decourty L, Doyen A, Malabat C, Frachon E, Rispal D, Séraphin B, Feuerbach F, Jacquier A & Saveanu C*. Long open reading frame transcripts escape nonsense-mediated mRNA decay in yeast. Cell Rep (2014), Feb 27;;6(4):593-8. PMID: 24529707
A multiplexed assay for over 800 NMD reporters identified coding sequence length as a major determinant of mRNA stability.